SpecRNA-QA: Why Spectral Graph Features See What Local Metrics Miss
RNA 3D structure, quality assessment, spectral graph theory, graph Laplacian, CASP16, bioinformatics, structural biology
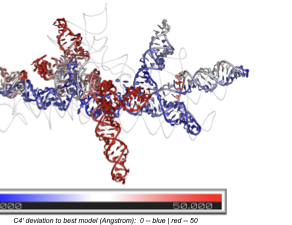
The Problem: Local Correctness, Global Failure
Predicting RNA 3D structure is one of the open frontiers of structural biology. Tools like AlphaFold3 and RoseTTAFold2NA now generate thousands of candidate structures — but how do you know which ones are right?
Existing quality assessment (QA) methods evaluate local atomic contacts: bond angles, clash scores, pairwise distance distributions. These work well when errors are local. But they fail catastrophically in a common and important failure mode: the local structure is correct, but entire domains are misplaced. A helix can be perfectly folded yet docked into the wrong pocket. Every bond angle checks out; the global topology is wrong.
This is exactly the regime where RNA QA matters most — large, multi-domain structures where the combinatorial space of domain arrangements dwarfs the local conformational space.
The Insight: Global Topology Lives in the Spectrum
A 3D molecular structure is, at its core, a graph: nucleotides are nodes, spatial contacts are edges. The spectrum of the graph Laplacian — the eigenvalues of \(\mathcal{L} = I - D^{-1/2}WD^{-1/2}\) — encodes the global connectivity pattern in a way that is invariant to rotation, translation, and node relabeling.
The key mathematical facts:
- The number of zero eigenvalues equals the number of connected components
- The Fiedler value \(\lambda_1\) measures how easily the graph can be bisected — a proxy for global compactness
- The spectral gap \(\lambda_1 - \lambda_0\) quantifies community separation
- Heat-kernel traces \(Z(t) = \sum_k e^{-\lambda_k t}\) capture multi-scale diffusion: small \(t\) probes local geometry, large \(t\) probes global topology
- The participation ratio \(\mathrm{PR}(\boldsymbol{\lambda}) = (\sum_k \lambda_k)^2 / \sum_k \lambda_k^2\) measures effective spectral dimensionality
A misplaced domain changes the large-\(t\) heat-kernel trace (disrupted long-range diffusion) while leaving the small-\(t\) trace nearly intact (local contacts are fine). This is precisely the information that local metrics cannot access.
The Method
SpecRNA-QA builds on this insight with a practical pipeline:
Multi-scale contact graphs: Construct contact networks at multiple distance thresholds (8Å, 10Å, 12Å, 15Å), capturing different spatial resolutions of the RNA architecture.
Spectral feature extraction: From each graph’s normalized Laplacian, extract ~312 features:
- Eigenvalue statistics (mean, variance, skewness, kurtosis of \(\{\lambda_k\}\))
- Heat-kernel traces at multiple diffusion times \(Z(t)\) for \(t \in \{0.1, 0.5, 1, 2, 5, 10\}\)
- Participation ratios and effective rank measures
- Spectral gap and algebraic connectivity
- Normalized Laplacian entropy \(H = -\sum_k \hat{\lambda}_k \log \hat{\lambda}_k\)
Learning-to-rank: An XGBRanker model trained to rank structures by quality within each target, using spectral features as input.
The entire pipeline runs on CPU — no GPU required. Processing time: 15 ms for a 100-nucleotide structure, ~4.2 seconds for 800 nucleotides.
Results
CASP16 Benchmark
On the CASP16 RNA structure prediction assessment (42 targets, 7,368 models):
| Method | Median Spearman \(\rho\) | \(p\)-value vs. SpecRNA-QA |
|---|---|---|
| SpecRNA-QA (supervised) | 0.689 | — |
| Geometry baselines | 0.465 | \(1.2 \times 10^{-10}\) |
Where It Matters Most: Large RNAs
The advantage is most pronounced for large RNA structures (>200 nucleotides), where the performance gap reaches +0.233 in Spearman correlation. This is the regime where domain-level misplacements dominate — and where spectral features shine.
For small RNAs (<100 nt), local metrics are often sufficient because there are few domains to misplace. The spectral advantage grows with structural complexity, exactly as the theory predicts.
Most Discriminative Features
Feature importance analysis reveals that the top-ranked features are heat-kernel traces at intermediate-to-large diffusion times — precisely the features that probe multi-scale and global transport geometry on the contact network. Local eigenvalue statistics (which probe small-scale structure) rank lower.
This confirms the theoretical motivation: the spectral approach works because it accesses the global information that local methods cannot reach.
Connection to the Broader Spectral Program
SpecRNA-QA is part of a broader research program applying spectral graph theory to structural biology:
- SpecRNA-QA (RNA): Multi-scale Laplacian spectra for RNA 3D quality assessment → under review at Briefings in Bioinformatics
- Spectral Coherence Index (Proteins): Participation-ratio effective rank of inter-model distance-variance matrices for protein ensemble QA, achieving AUC-ROC 0.973 on 110 NMR ensembles → under review at IEEE JBHI (arXiv:2603.25880)
Both methods share a design principle: model-free spectral features that are invariant to coordinate systems and capture global structural properties that local metrics miss.
Try It
SpecRNA-QA is open source and easy to use:
git clone https://github.com/yudabitrends/specrnaq
cd specrnaq
pip install -e .
specrnaq predict --input structures/ --output scores.csvPython 3.10+, CPU-only, no external dependencies beyond standard scientific Python.
Papers
Ying Zhu, Huaiwen Zhang, Vince D. Calhoun†, Yuda Bi†. Spectral Graph Features Capture Global Topology for Reference-free RNA 3D Structure Quality Assessment. Under review at Briefings in Bioinformatics.
Yuda Bi, Huaiwen Zhang, Jingnan Sun, Vince D. Calhoun. Spectral Coherence Index: A Model-Free Metric for Protein Structural Ensemble Quality Assessment. Under review at IEEE JBHI. arXiv:2603.25880